Center Update: Improving Freeze Tolerance in Turfgrasses
go.ncsu.edu/readext?759734
en Español / em Português
El inglés es el idioma de control de esta página. En la medida en que haya algún conflicto entre la traducción al inglés y la traducción, el inglés prevalece.
Al hacer clic en el enlace de traducción se activa un servicio de traducción gratuito para convertir la página al español. Al igual que con cualquier traducción por Internet, la conversión no es sensible al contexto y puede que no traduzca el texto en su significado original. NC State Extension no garantiza la exactitud del texto traducido. Por favor, tenga en cuenta que algunas aplicaciones y/o servicios pueden no funcionar como se espera cuando se traducen.
Português
Inglês é o idioma de controle desta página. Na medida que haja algum conflito entre o texto original em Inglês e a tradução, o Inglês prevalece.
Ao clicar no link de tradução, um serviço gratuito de tradução será ativado para converter a página para o Português. Como em qualquer tradução pela internet, a conversão não é sensivel ao contexto e pode não ocorrer a tradução para o significado orginal. O serviço de Extensão da Carolina do Norte (NC State Extension) não garante a exatidão do texto traduzido. Por favor, observe que algumas funções ou serviços podem não funcionar como esperado após a tradução.
English
English is the controlling language of this page. To the extent there is any conflict between the English text and the translation, English controls.
Clicking on the translation link activates a free translation service to convert the page to Spanish. As with any Internet translation, the conversion is not context-sensitive and may not translate the text to its original meaning. NC State Extension does not guarantee the accuracy of the translated text. Please note that some applications and/or services may not function as expected when translated.
Collapse ▲Improving freeze tolerance in warm-season grasses
PI: Susana R. Milla-Lewis; Student: Jessica M. Brown
Introduction
In comparison with other warm-season grasses, Zoysiagrass (Zoysia spp.) has moderate drought and shade tolerances. However, one factor limiting the widespread use of zoysiagrass is a relative lack of freeze tolerance, especially when compared to cool-season grasses. (Patton and Reicher, 2007). ‘Meyer’ was released in the 1950s for its excellent freezing tolerance and is still considered standard for this trait. Somewhat limited progress has been made in the development of more freeze tolerant zoysiagrass cultivars. This is in part due to the complexity of the trait, insufficient genetic knowledge of freeze tolerance components, and lack of efficient selection criteria. A project was initiated in 2013 to improve the efficiency of selection for freeze tolerance in zoysiagrass by identifying genomic regions controlling the trait and using DNA markers associated with them for selection.
Methods
Freeze tolerant ‘Meyer’ was crossed to freeze-susceptible ‘Victoria’ to generate a population that was segregating for the trait. A total of 175 individuals were evaluated for freeze tolerance under controlled environmental conditions -in order to avoid escapes- and including acclimation treatments for a better assessment of true freeze response. Plants were vegetatively propagated in cone-tainers (2.5×12 cm) using a single stolon and allowed to establish for at least 6 weeks in the greenhouse. After establishment, plants were cold acclimated (CA) for 4 weeks in a growth chamber at 8/2°C day/night cycles with a 10-h photoperiod. Non-acclimated (NA) plants remained in the greenhouse at 27°C during this 4 week period. Immediately before freezing, NA and CA plants were randomized together. Plants were arranged in freezers in a randomized complete block design with two replications. Based on reported LT50s (Pruitt et al. 2016), freezing temperatures were -9°C, and -11°C. After freezing, plants were thawed overnight and returned to the greenhouse. Surviving green tissue (SGT) and regrowth (RG) ratings were collected one day after freezing and then once a week for 6 weeks post-freeze. SGT, which represents the severity of injury a plant sustains during freezing, was rated on a scale of 0-5 with 0=all tissue and nodes were brown, and 1=0-25%, 2=25-50%, 3=50-75%, 4=75-99%, and 5=100% green tissue and nodes. RG, based on the percent of new green leaves and nodes, was also rated on the same 0-5 scale with 0=0% and 5=100% new tissue. All ratings were converted to percentages for data reporting and interpretation.
Freezing data and our existing linkage map were used for genetic analysis using composite interval mapping via JMP Genomics 7 (SAS Institute; Cary, NC). Single marker analysis was first performed to identify regions strongly correlated with freezing tolerance. Then, interval mapping (IM) and composite interval mapping (CIM) procedures were used to identify genomic regions associated with freeze tolerance. These and those identified by Holloway et al. (2018) based on field winter survival were compared to identify overlap. Sequence analysis in those overlapping regions was performed in order to scan them for presence of genes that had been previously identified as having a role in freeze or stress tolerance in other species.
Results and Discussion
Freezing Tests
In controlled laboratory freezing tests, the effects of genotype and cold acclimation treatment were found to be significant for both SGT and RG (Table 1). Compared to NA, freezing tolerance under CA was markedly improved for both traits. For CA, SGT was increased by 14.4-fold (from 0.067 to 0.967), 2.3-fold (from 0.333 to 0.767) and 3.8-fold (from 0.210 to 0.796) in Meyer, Victoria and the progeny, respectively. Meanwhile, RG was improved by 13.4-fold (from 0.067 to 0.900), 2.2-fold (from 0.300 to 0.667) and 3.5-fold (from 0.202 to 0.697) in Meyer, Victoria and the progeny, respectively (Table 1).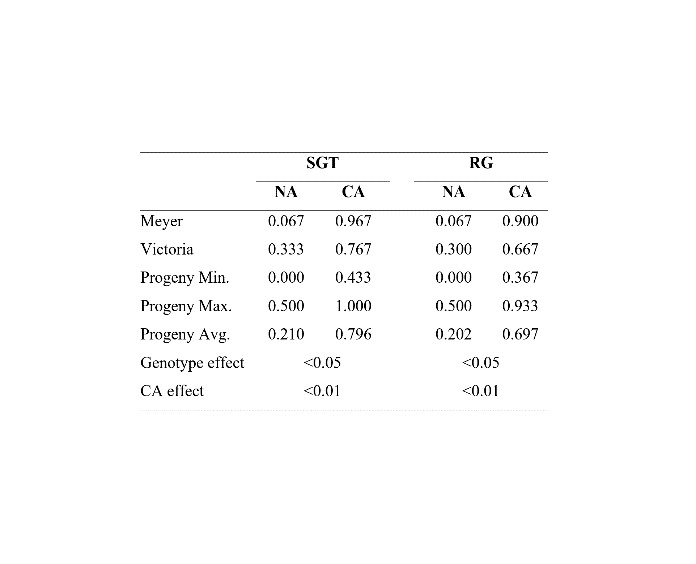
Among the parents, Meyer performed better under CA for both SGT and RG; For both SGT and RG, progeny with better performance than both parents was observed under all environmental conditions.
Genomic Regions Associated with Freeze Tolerance
A total of thirty-nine regions were identified that explained between 6.4 and 12.2% of the phenotypic variance. Among these, 21 of them were identified only under CA, 15 under NA, while 3 were found to be significant under both CA and NA conditions. There were nine regions specifically associated with SGT, eight linked to RG, and 22 QTL were found to be significant for both SGT and RG. These regions of interest were compared to those associated with field winter injury (WI) in Holloway et al (2018). Eleven regions of overlap were identified between both studies (Figure 1). Among these, four were found to be shared between WI and RG, and four were common between WI and SGT. Of particular interest were the three regions that overlapped among all traits.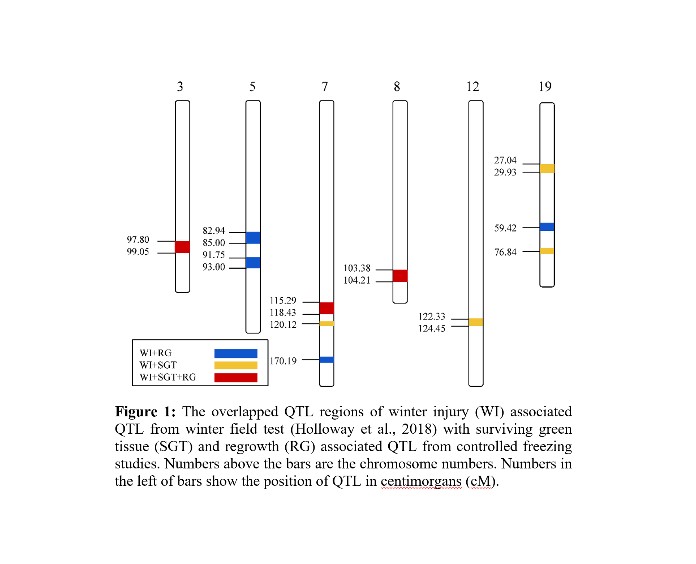
Sequence annotation in regions of interest
Sequences within the regions of interest were used to search for homology with previously identified genes related to abiotic stress tolerance, particularly freeze tolerance. Several abiotic response proteins were found underlying these overlapping regions, such as transcription factors, cell wall modification related proteins, and defense signal transduction related proteins. However, further studies will need to be conducted to determine gene expression in response to cold acclimation and freezing tolerance in zoysiagrass. Following further validation, these genomic regions and their associated markers have the potential to be utilized in future breeding efforts for the development of a broader pool of zoysiagrass cultivars capable of surviving in cooler climates.
A manuscript summarizing this work is currently under review for publication in Molecular Breeding.
Holloway, H.M.P., X. Yu, J.C. Dunne, B.M. Schwartz, A.J. Patton, C. Arellano and S.R. Milla-Lewis. 2018. A NP-based high-density linkage map of zoysiagrass (Zoysia japonica Steud.) and its use for the identification of QTL associated with winter hardiness. Molecular Breeding, 38(1). doi:10.1007/s11032-017-0763-0
Patton, A.J. and Z.J. Reicher. 2007. Zoysiagrass species and genotypes differ in their winter injury and freeze tolerance. Crop Science 47:1619-1627.
Pruitt, M.H. 2016. Genomic approaches for improving freeze tolerance in zoysiagrass. MS Thesis. North Carolina State University, Raleigh, NC.


